Search help
Our search options allow users to search all of our active gene symbol reports and gene group reports quickly and with ease. The search server utilises Apache Solr which offers a powerful full-text search, hit highlighting and faceted searching. Faceted searching allows you to search for a particular keyword and then filter the results by record/page type, locus group and locus type.
The search form can be found within the masthead of each page. Enter a search term within the search input box and click on the spy glass icon; the default option searches the whole site including gene symbol reports and gene groups. Users can also click on the drop down to the left of the input box and search only specific areas of the site (i.e symbols or gene groups).
Basic search
The simplest way to search is to type a query word/ID into the input box within the masthead and click on the spy glass while the dropdown specifies "Search all". The default option is a full-text search over all the indexes and fields. If reports are found containing the keyword/ID they are displayed in order of relevance (for more information see indexed fields for each search type). On the left hand side of the results page the filter options (where available) are shown with the numbers of reports associated with each facet. If the results include gene symbols then clicking on the facet "Gene" will filter the results by this type and will change the faceting to display the locus groups and types that are relevant to the search results; enabling further filtering by locus type. Users can also change the default number of results per page from 10 up to 200.
The results display specific fields from within the search index which differ depending on the document type. The first line of each result contains the gene symbol and the gene name if the result is a gene symbol report or the group name if it's a gene group report. The second row will show the type of the indexed document (i.e gene, or group) and will also contain some of the important fields to help identify the hit. The third row reports the field the keyword/ID matches, so if the keyword matched an approved symbol within a gene symbol report the third row would say "Matches: Gene symbol etc" as seen below in figure 1.
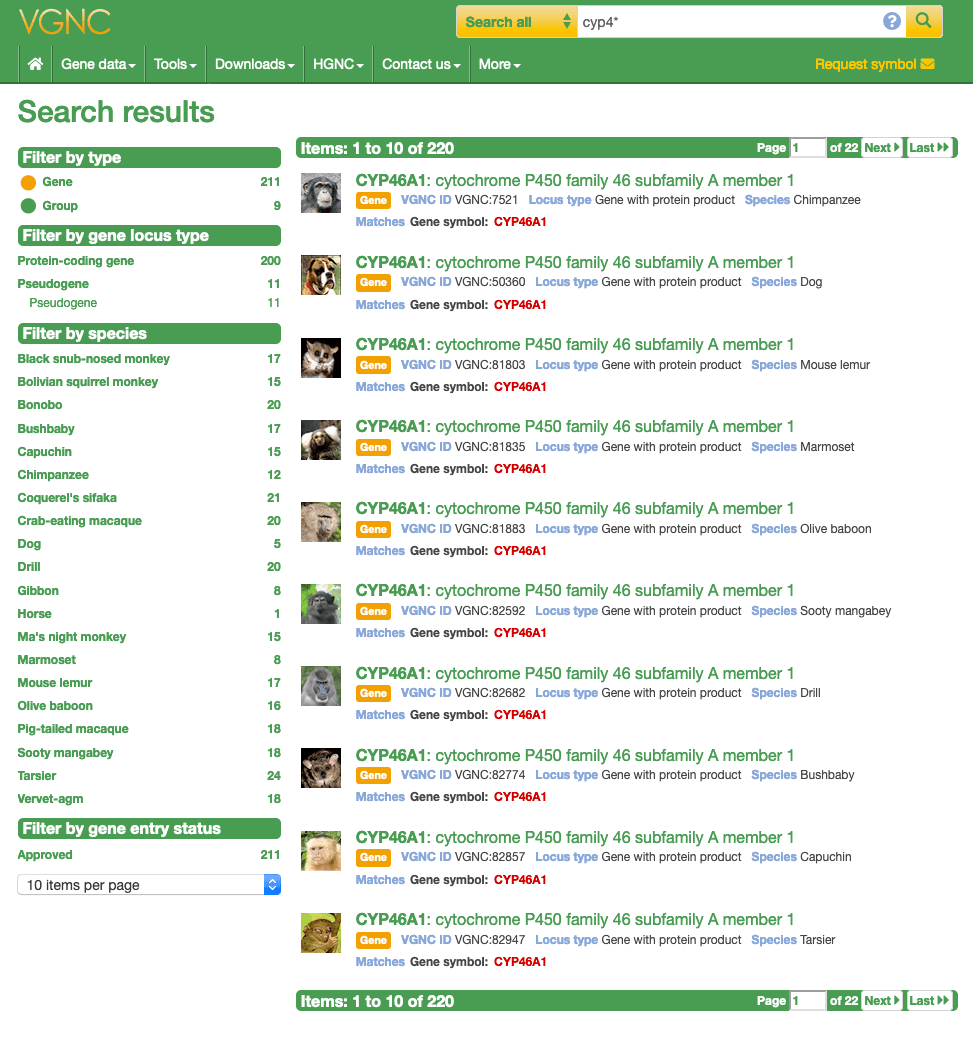
Advanced search
The search application allows users to make advanced queries using the search box. In this section we describe how to specify the search type, use wildcards, logic operators and specify indexed fields.
Search types
Instead of searching everything, users can select to search only for gene symbol reports, or for gene groups only by selecting these options from the dropdown next to the search input box within the masthead.
Wildcard search
Sometimes it may be useful to match records based on a query pattern rather than a keyword or ID. Our search allows users to use wildcard operators with an asterisk (*) to stand in place for one or more characters and a question mark (?) to stand in place for a single character substitution. Multple wildcards can be used in the same query, for instance searching for AB?1* will find the symbols ABI1, ABL1, ABT1.
Logic operators
By default the search application uses the logical operator OR, so inputing BRAF RNA into the search box actually equates to BRAF OR RNA so some of the results will contain BRAF and some will contain RNA and others may contain both. Sometime however this isn't what a user wants to search. Lets say a user wants to find reports that contain both BRAF and RNA in the same report. Typing in BRAF RNA will unfortunately return over 1000 hits. By changing the search query to BRAF AND RNA the results returned are more pertinent and reduces the number of hits returned. Alternatively asking for reports that do not contain a keyword/ID may be preferable therefore users may use NOT or - within term such as BRAF NOT RNA or BRAF -RNA.
Phrases
As discussed above the default operator is OR, so if a search using the term protein arginine methyltransferase 1 was used the actual search term will be protein OR arginine OR methyltransferase OR 1. This can be addressed by simply quoting the query so that the search knows to treat the quoted block as one term like "protein arginine methyltransferase 1".
Indexed fields
Users may search within a specific indexed field by using the information seen in the indexed fields section below using a very basic notation. To specify an index the user need only to type in to the search field the indexed field key followed immediately with a colon (:) and then the query eg entrez_id:463781. If the query is not an ID or a keyword "phrases" can be used after the colon eg gene_prev_name:"cysteine rich angiogenic inducer 61".
Indexed fields
"Search all" fields
- ensembl_gene_id
- Ensembl gene ID. Found within the "Gene resources" section of the gene symbol report e.g. ENSFCAG00000010251
- group_alias
- Other names used to refer to this group as seen within the gene group reports under the "Also known as" field.
- group_id
- The VGNC gene group ID which can be seen in the URL for a gene group page after the word "group" e.g. 46
- group_name
- The gene group name as set by the VGNC and seen at the top of the gene group reports e.g. Cytochrome P450 family 5
- gene_alias_name
- Other names used to refer to this gene as seen in the "Alias names" field in the gene symbol report e.g. cytochrome P450, family 5, subfamily A member 1
- gene_alias_symbol
- Other symbols used to refer to this gene as seen in the "Alias symbols" field in the gene symbol report e.g. CYP5A1
- gene_curator_notes
- Contains additional information related to an entry that has been manually added by an HGNC curator
- gene_name
- VGNC approved name for the gene. Equates to the "Approved name" field within the gene symbol report e.g. zinc finger protein 536
- gene_prev_name
- Gene names previously approved by the VGNC for this gene. Equates to the "Previous names" field within the gene symbol report e.g. hyaluronoglucosaminidase 1
- gene_prev_symbol
- Gene symbols previously approved by the VGNC for this gene. Equates to the "Previous symbols" field within the gene symbol report e.g. SEPT1
- gene_symbol
- The VGNC approved gene symbol. Equates to the "Approved symbol" field within the gene symbol report e.g. KLF4
- locus_group
- A group name for a set of related locus types as defined by the VGNC e.g. pseudogene.
- locus_type
- The locus type as set by the VGNC e.g.gene with protein product
- ncbi_gene_id
- NCBI gene ID. Found within the "Gene resources" section of the gene symbol report e.g. 101082021
- root_symbol
- The common root gene symbol associated to a group if a common root symbol exists.
- symbol_status
- Status of the symbol report, which can be either "Approved" or "Entry Withdrawn" e.g. Approved
- uniprot_id
- UniProt protein accession. Found within the "Protein resource" section of the gene symbol report e.g. A5PJU9
- vgnc_id
- VGNC ID. A unique ID created by the VGNC for every approved symbol e.g. VGNC:1097
"Search Genes" fields
- alias_name
- Other names used to refer to this gene as seen in the "Alias names" field in the gene symbol report e.g. cytochrome P450, family 5, subfamily A member 1
- alias_symbol
- Other symbols used to refer to this geneas seen in the "Alias symbols" field in the gene symbol report e.g. CYP5A1
- ensembl_gene_id
- Ensembl gene ID. Found within the "Gene resources" section of the gene symbol report e.g. ENSFCAG00000010251
- locus_group
- A group name for a set of related locus types as defined by the VGNC e.g. pseudogene
- locus_type
- The locus type as set by the VGNC e.g. gene with protein product
- name
- VGNC approved name for the gene. Equates to the "Approved name" field within the gene symbol report e.g. zinc finger protein 536
- ncbi_gene_id
- NCBI gene ID. Found within the "Gene resources" section of the gene symbol report e.g. 101082021
- prev_name
- Gene names previously approved by the VGNC for this gene. Equates to the "Previous names" field within the gene symbol report. e.g. hyaluronoglucosaminidase 1
- prev_symbol
- Symbols previously approved by the VGNC for this gene. Equates to the "Previous symbols" field within the gene symbol report e.g. SEPT1
- status
- VGNC status for gene symbol reports the values of which will either be "Approved" or "Entry Withdrawn" e.g. Approved
- symbol
- The VGNC approved gene symbol. Equates to the "Approved symbol" field within the gene symbol report e.g. KLF4
- uniprot_ids
- UniProt protein accession. Found within the "Protein resource" section of the gene symbol report e.g. A5PJU9
- vgnc_id
- VGNC ID. A unique ID created by the VGNC for every approved symbol e.g. VGNC:1097
"Search Gene groups" fields
- group_alias
- Other names used to refer to this group as seen within the gene group reports under the "Also known as" field.
- group_id
- The VGNC gene group ID which can be seen in the URL for a gene group page after the word "group" e.g. 46
- group_name
- The gene group name as set by the VGNC and seen at the top of the gene group reports e.g. Cytochrome P450 family 5
- vgnc_id
- VGNC ID. A unique ID created by the VGNC for every approved symbol e.g. VGNC:42517
- root_symbol
- The common root gene symbol associated to a group if a common root symbol exists e.g. CYP